On 10–12 February, PHUSE held the inaugural Nonclinical Advance Event, themed ‘The Power Of Nonclinical Data: Enabling NAMs, Predictive Modelling And Machine Learning For Translational Safety’.
As PHUSE’s first multi-day nonclinical event, it successfully delivered a focused virtual forum exploring how data is transforming translational safety.
Over three days, 487 attendees from 99 unique companies across 22 countries joined virtually. Sessions explored the role of nonclinical data in enabling NAMs, predictive modelling and machine learning, with practical insights for industry application.
All presentation slides and recordings are available on the PHUSE website. Continue reading for reflections on each day.
Day 1: New Approach Methodologies for Nonclinical Safety Assessment
Day 1 of the Nonclinical Advance Event focused on how new approach methodologies (NAMs) are moving from policy discussion into practical regulatory implementation. Across regulatory, industry and standards perspectives, a clear message emerged: innovation must be supported by aligned data structures, consistent terminology and collaborative frameworks to build regulatory confidence.
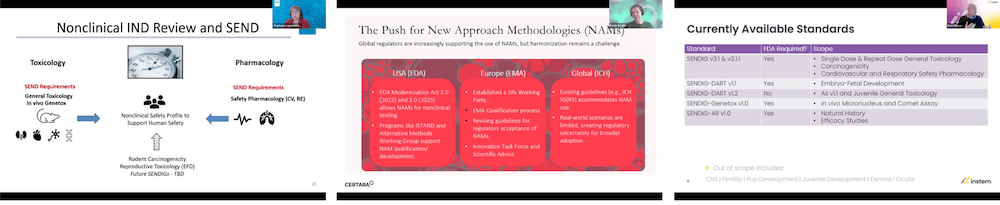
The day opened with insights from the FDA’s CDER, where Nakissa Sadrieh and Stephanie Leuenroth-Quinn outlined how legislative updates, including FDORA, had enabled broader use of NAMs in regulatory submissions. They described how CDER had operationalised NAM review through structured processes, including a NAM Integrated Review Team, and clarified the distinction between qualification and validation. SEND was highlighted as a key tool for improving review efficiency, while challenges remain around standardised nomenclature and the diversity of in vitro data.
Veronique Francois-Newton then introduced the Pistoia Alliance’s mission to lower barriers to innovation through pre-competitive collaboration. Her session emphasised that harmonised, AI-ready data structures are foundational to interoperability across researchers, toxicologists, CROs and regulators.
Chris Butler presented the In Vitro Pharmacology (IVP) Project, which addressed the lack of consistent standards for in vitro assay data. By developing a shared ontology, an assay registration system, and a structured submission pathway, the project aimed to accelerate regulatory review and improve data reuse.
Kevin Snyder expanded on this theme with a proposed In Vitro NAMs data model built around assay metadata, study design and study results. Designed to reduce fragmentation and support AI-ready integration, the model is being developed in collaboration with the FDA, CDISC and FNIH to ensure regulatory sustainability.
Closing the session, Marc Ellison explored the absence of harmonised standards for in vivo efficacy data and proposed expanding SEND beyond toxicology. Planned new domains for disease models and efficacy endpoints aim to bring greater consistency and unlock broader data reuse.
In the breakout discussions, participants acknowledged both opportunity and complexity. Extending SEND into the NAMs space will require balancing flexibility with structure.
Alignment on terminology was identified as an essential first step, with the suggestion of beginning with a focused set of NAM types before scaling more broadly. Standards were widely viewed as enablers of automation and predictive modelling, but success will depend on early collaboration between scientists, data standards experts, regulators and industry.
By the end of Day 1, the direction was clear: advancing NAMs is not only a scientific challenge, but a data and alignment challenge. Thoughtful standardisation, cross-sector collaboration and practical pilot initiatives will be key to translating potential into routine regulatory practice.

Day 2: Nonclinical Predictive Modelling & Machine Learning
Day 2 of the Nonclinical Advance Event shifted from standards to application, exploring how predictive modelling and machine learning are transforming nonclinical data into translational insights. Where Day 1 focused on alignment and structure, Day 2 examined how structured data can be actively leveraged to support earlier and more informed decision-making.
The day opened with Panteleimon Mavroudis presenting hybrid and hierarchical machine learning frameworks designed to predict pharmacokinetics directly from molecular structure. In preclinical species, the models predicted plasma exposure within a two- to threefold range of observed values. Extension to humans required substantial data curation, including the development of an external curated PK database. The session demonstrated both the potential of ML-enabled translational modelling and the importance of aligning model choice with the scientific question and data assumptions.
Kevin Snyder then outlined the PHUSE Predictive Modelling Project, a collaborative initiative focused on applying predictive analytics to CDISC SEND-formatted toxicology data. Through a structured three-phase roadmap, the project is sharing existing approaches, cross-testing methods across organisations and working towards reusable modelling tools. Current efforts include endpoint normalisation, SEND-based scoring and benchmarking, with longer-term goals of improving signal detection and supporting translation of nonclinical findings to human safety outcomes.
The final presentation from Lilliam Rosario and William Houser broadened the lens to translational safety and collaborative toxicology data sharing. They described an inflexion point in safety science, where harmonised datasets, NAMs and predictive analytics can strengthen the connection between nonclinical observations and clinical outcomes. Improved terminology alignment and interoperability were highlighted as essential enablers of this vision.
The breakout discussions reflected both ambition and pragmatism. Participants emphasised the importance of digitising expert conclusions from study reports, capturing not only raw SEND outputs but also structured interpretations to enhance model training. Data sharing emerged as a significant challenge, with intellectual property constraints limiting openness despite strong internal modelling activity across organisations. Collaborative platforms such as PHUSE were seen as essential in providing governance frameworks that support responsible exchange. Alternatives, including code sharing and federated learning, were discussed as potential pathways.
Bias and representativeness were also key themes. Models trained predominantly on compounds that reach clinical development may overlook more informative toxicity signals found in failed programmes. Addressing dataset breadth and actively interrogating bias were recognised as critical next steps.
Finally, participants stressed the need to think beyond submission requirements. If predictive analytics are to scale, study design, metadata capture and structured recording of results must evolve accordingly. Education initiatives to bridge SEND expertise and modelling needs were suggested as practical ways to strengthen this alignment.
By the close of Day 2, the message was clear: predictive modelling in nonclinical science is no longer theoretical. Its continued impact will depend on high-quality data, thoughtful standardisation and sustained cross-sector collaboration.
Day 3: PHUSE Project Contributions & Developments
Day 3 of the Nonclinical Advance Event shifted from ‘what’s possible’ to ‘what needs to work well’ – focusing on practical SEND implementation, quality, and how the community can better support consistent data use beyond submission.
The day opened with highlights from the SEND Survey 2026, which reinforced recurring pain points such as software issues, offline data formats, and uneven adoption of guidance, alongside continued discussion of tumour.xpt and non-standard implementations. A key message was the need for clearer communication and stronger shared resources for the community.
The Tumor.xpt Conformance Group then outlined progress on developing clearer conformance rules and validation approaches for tumour data submissions, with the aim of reducing manual review effort and improving consistency. This was followed by a 10-year milestone update on the nSDRG, emphasising the ongoing value of structured context and reviewer guidance alongside SEND datasets.
Updates from the SEND Coding Bootcamp highlighted momentum in building practical programming and data-handling skills across sponsors, CROs and regulators, with a growing focus on applied learning through scripts, demos and office hours. The day closed with a look at Virtual Control Groups (VCGs) and the role SEND can play in supporting them, while acknowledging the ongoing need for standardised metadata and cross-industry alignment.
In the Day 3 breakout, participants reflected that the discussion ended early but generated strong, actionable ideas for future PHUSE work. One group proposed developing best practices for NAMs using a TIG-style approach, with a focus on flexibility and scalability while standards mature. A practical suggestion was to reduce barriers for scientists by creating an Excel-based data capture template paired with open-source converters to generate JSON/XPT outputs (and potentially Define-style metadata), with the idea of building this as a collaborative ‘capstone-style’ project through the Coding Bootcamp.
The breakout also highlighted strong interest in using JSON datasets as a modern path for NAMs and explored how PHUSE could accelerate adoption through collaboration and shared tooling. Participants discussed potential support for SEND/SDTM integration work underway elsewhere, and several attendees flagged the value of reviving a searchable Q&A/wiki model (rather than relying on chatbots) given current limitations in LLM accuracy for SEND questions. Overall, the conversation reinforced that the community is ready to prioritise practical tools, shared learning, and stronger cross-disciplinary input to keep SEND fit for expanding use cases.

